Biological physics of dense microbial populations
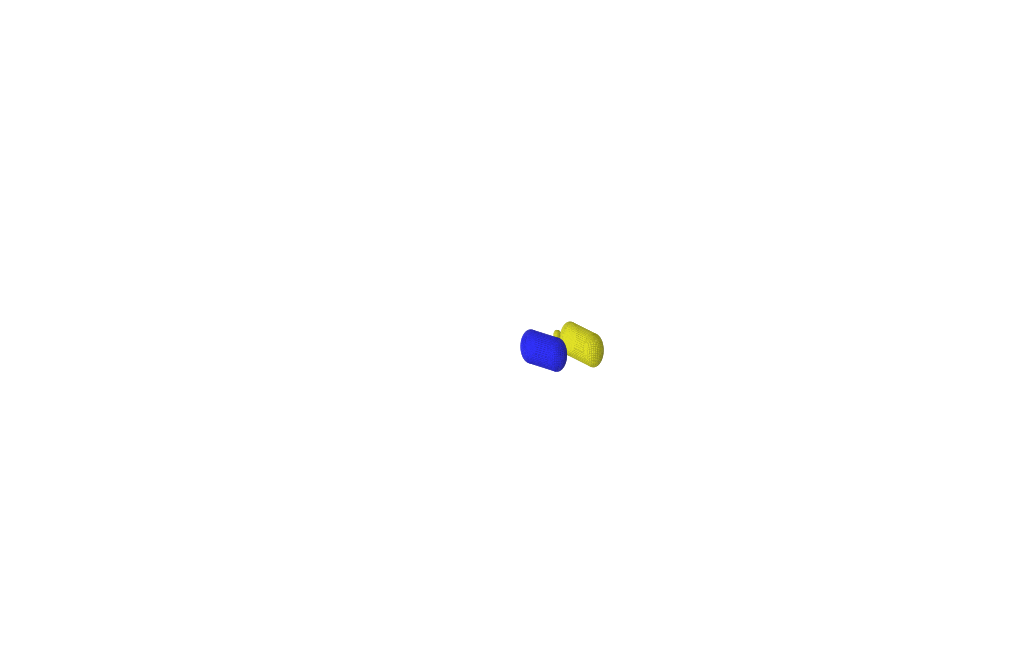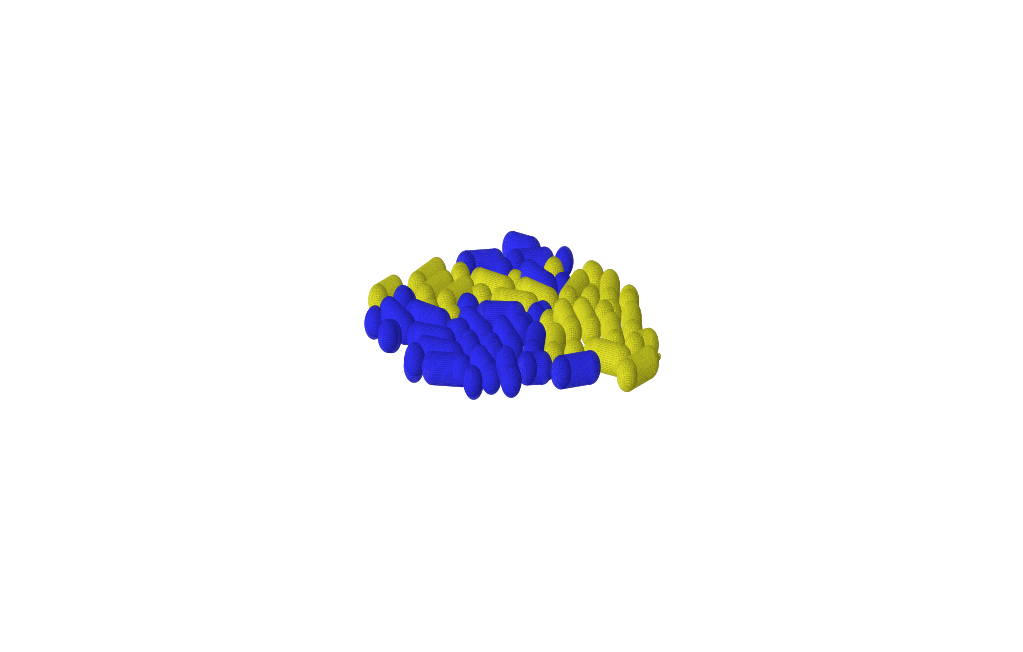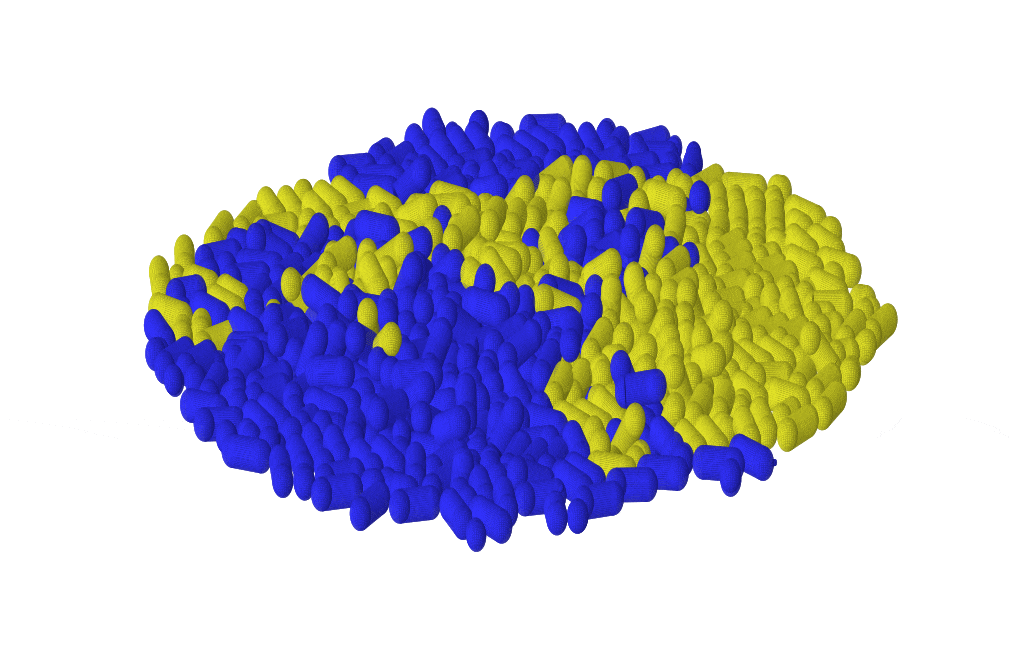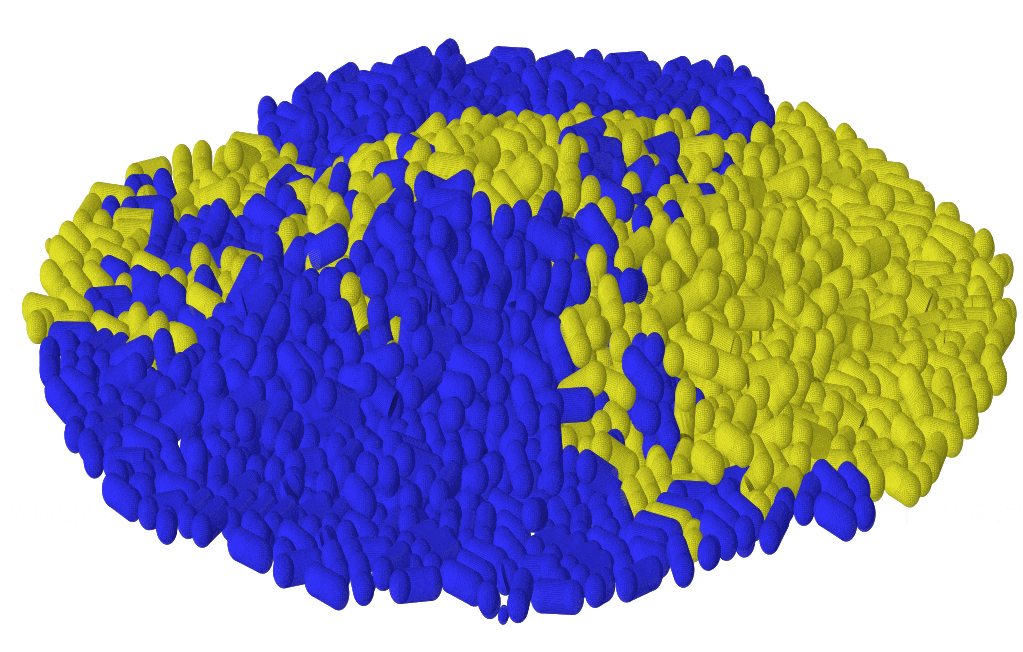
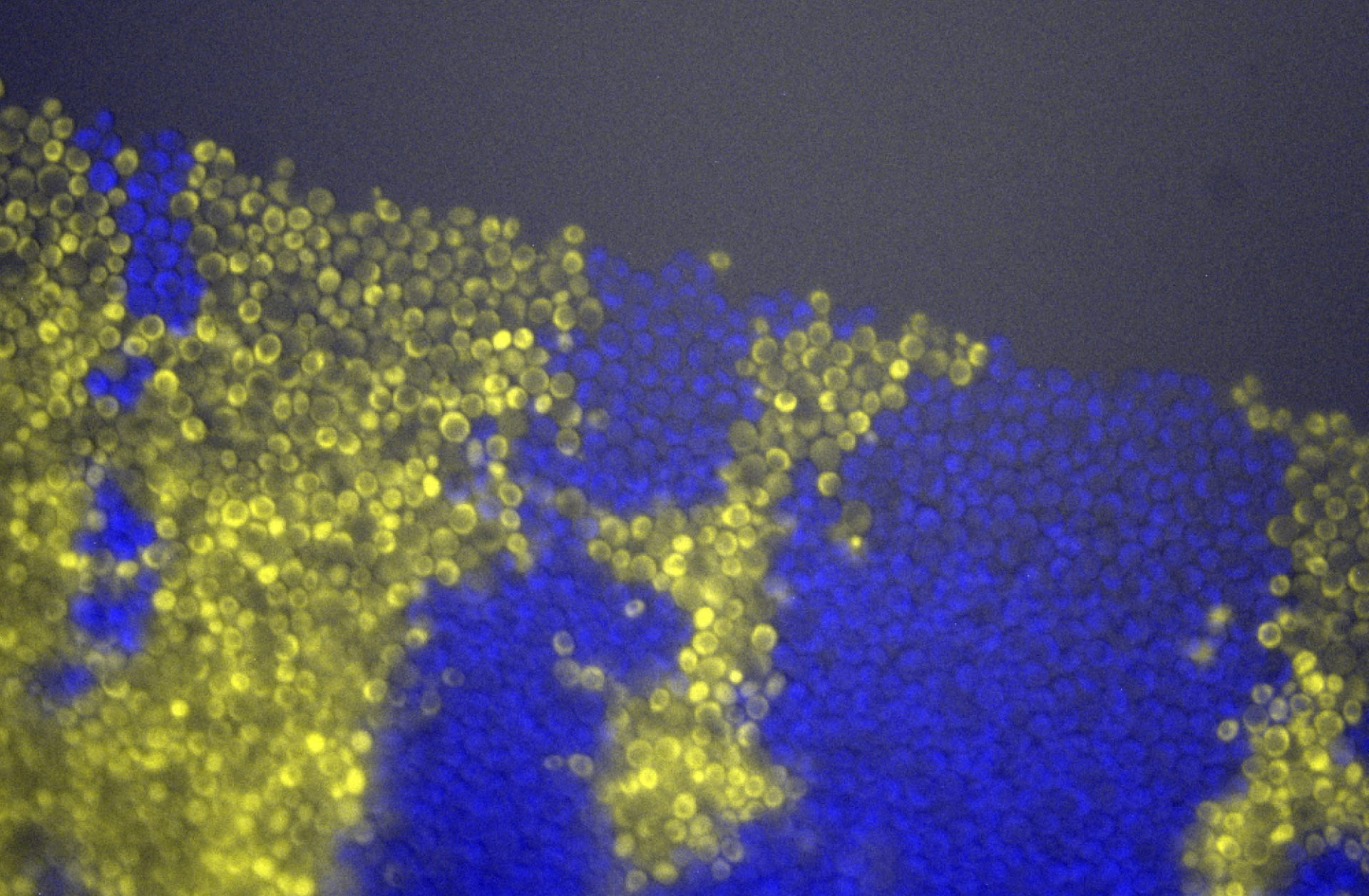
Microbes often assemble in dense populations in which proliferating individuals exert mechanical forces on the nearby cells.
Mechanical interactions among cells affect the competition between different genotypes in growing yeast colonies, causing the prolonged survival of slower-growing strains at the actively-growing frontier of the colony (forming thin, persistent filaments that survive for very long times), and causing faster-growing strains to increase their frequency more slowly than expected in the absence of mechanical interactions. Thus, mechanical interactions among growing, dividing cells reduce the power of natural selection in dense populations of cells, which may favor the maintenance of drug-resistant mutants in expanding microbial populations, and may explain the alleged neutrality of inter-clonal competition in growing tumors.
These effects also promote the survival of slower-growing strains and the maintenance of genetic diversity in colonies grown in time-varying environments in which the selective advantage of one strain over another alternate between different phases of the environment.
We investigate the role of mechanical interactions in shaping microbial population genetics via laboratory experiments with genetically engineered yeast and bacteria, mathematical modeling of microbial colonies as viscous, self-replicating fluids, and via numerical simulation of individual cells that grow, divide, and push each other.
Microbial warfare, cheating and cooperation
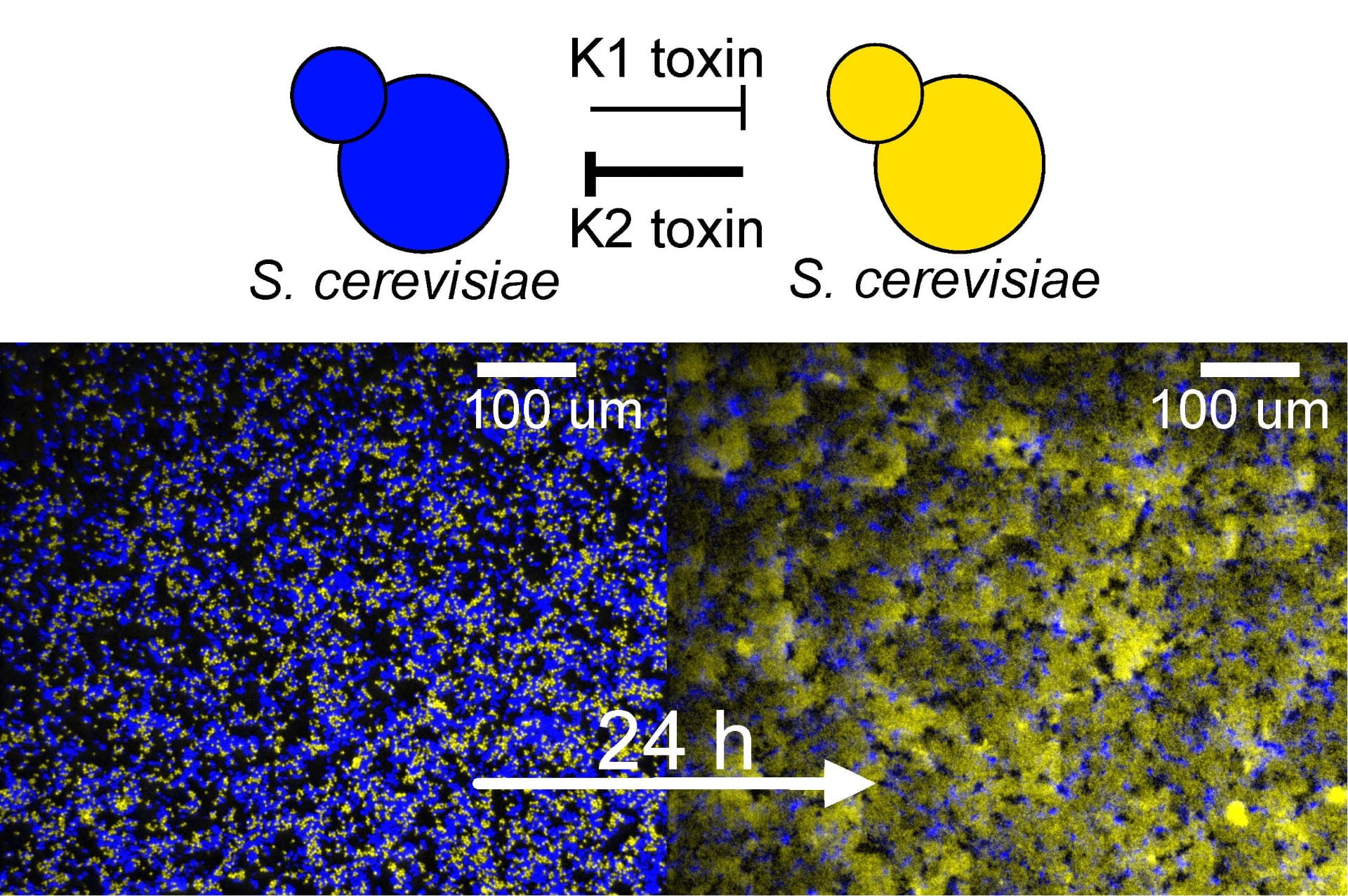
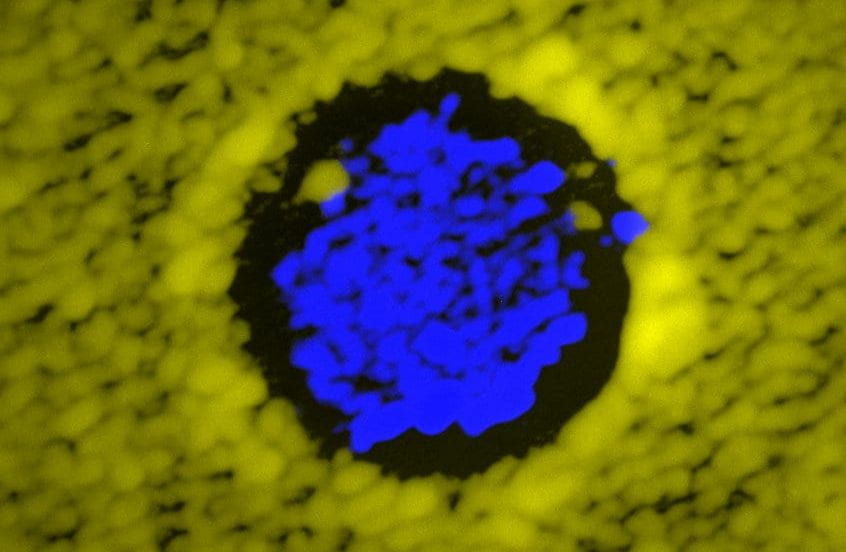
We investigate how microbial interactions such as cooperation, cheating and conflict affect the spatial assembly and expansion of microbial populations.
We genetically engineered strains of the budding yeast Saccharomyces cerevisiae to release toxins at experimentally tunable rates. With this system, we can experimentally investigate the spatial dynamics of microbial antagonism varying the strength of the interaction. This allows us to test experimental predictions of theoretical models of population dynamics with antagonistic interactions, refine mathematical models of microbial antagonism, and investigate experimentally the ecological and evolutionary dynamics of microbial warfare.
Using this experimental system, we showed that being a stronger antagonist is a necessary, but not a sufficient condition for successfully invading a landscape occupied by a resident, antagonist strain. In fact, a successful invasion also requires that the initial inoculum be larger than a critical size. We showed that mathematical models of microbial antagonism parametrized using well-mixed competitions between antagonist and sensitive strains can predict the dynamics of microbial warfare in both well-mixed and spatially-structured communities.
Furthermore, we observe that resistant cells and antagonists with reduced strength emerge in spatially-structured competitions between antagonist strains, suggesting that adaptive evolution can alter the dynamics of microbial antagonism, which is a research direction that we are currently pursuing.
Evolutionary dynamics of range expansions

Much of what we know about evolutionary dynamics comes from studies that used well-mixed populations. In well-mixed populations, evolution selects for traits that facilitate survival or favor reproduction. In spatially-expanding populations, however, evolution may also select for traits that enhance the rate of dispersal.
We study the evolutionary dynamics of range expansions through laboratory experiments with yeast and bacteria, and mathematical and numerical modeling.
Microbial community assembly
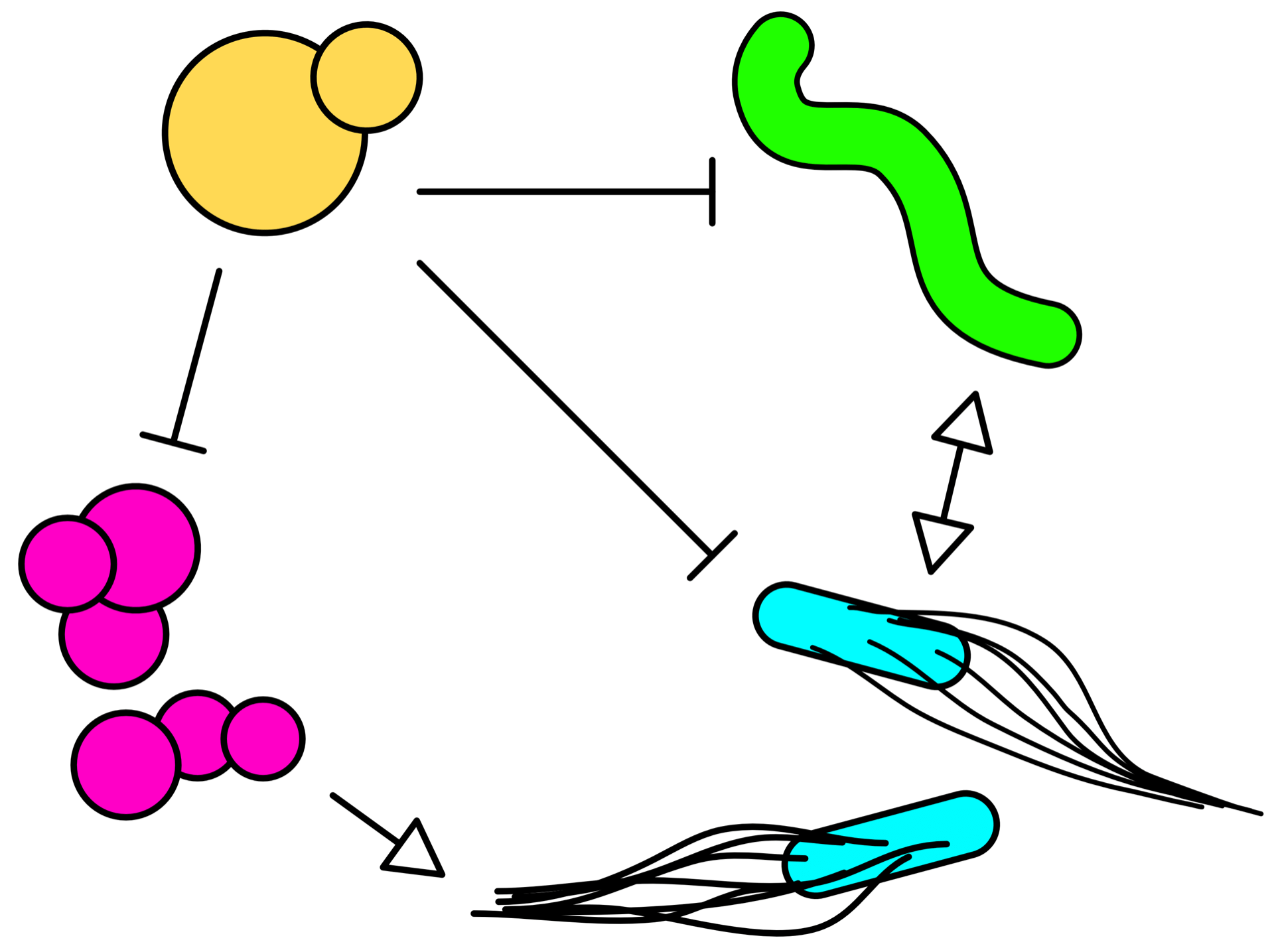
Which are the rules that govern community assembly? Can multiple species that compete for the same resources coexist in the same environment? How do interactions such as metabolites exchange and antagonism affect the composition and stability of microbial communities? We investigate these questions by combining theoretical models with laboratory experiments with genetically engineered microbes.
